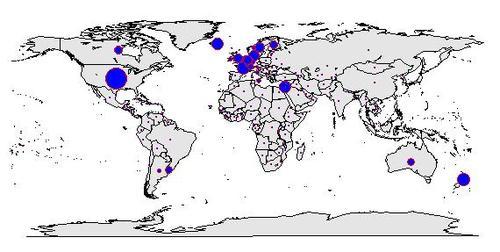
As mentioned on a previous post, I am interested in analysing if people’s ‘unhealthy’ lifestyle is associated to new cases of cancer diagnosed globally. The outcome variable I want to explore (at least for now), is the number of new cases of breast cancer in 100,000 female residents. I have this
data for 173 countries from 2002, as collected by ARC (International Agency for Research on Cancer).
So lets start studying breast cancer variable. Here is
some univariate plots I’ve made with R studio in order to understand breast cancer spread and distribution globally.
> range(gapCleaned$breastcancer)
[1] 3.9 101.1
Looking at all countries, breast cancer ranges from a minimum of 3.9 new cases to a maximum of 101.1 new cases per 100,000 female residents.
Let see more in details
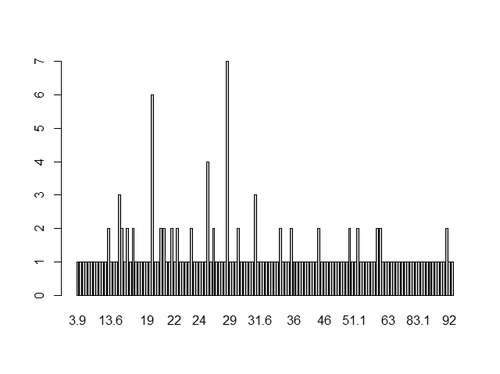
the distribution of breast cancer data, that is the frequency of occurrence of each value. We can visualize it through an histogram:
> hist(gapCleaned$breastcancer,10, main=”Breast cancer global distribution”, xlab=”# of new cases”, col=”blue”, border=”red”)
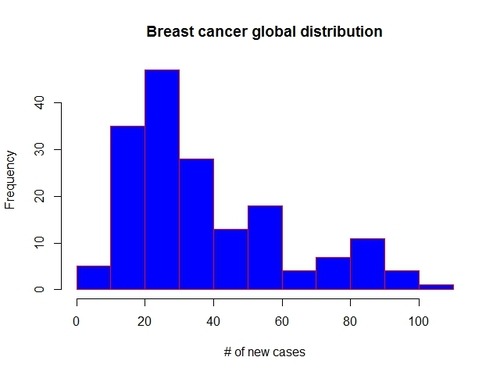
A couple of considerations on the shape of the distribution:
- it’s a unimodal distribution (there is only one peak: the mode), which means that most countries in our data set present a number of new cases of breast cancer between 20 and 30
- there are no outliers (no gaps in the data)
- the distribution looks asimmetric, right-skewed, as the there is a longer right tail. This implies that the mode is less than the median which in turn is less than the mean. The mean is larger than both median and mode, because it is influenced by the longer right tail of the data.
We can easily calculate the above parameters with summary function:
> summary(gapCleaned$breastcancer)
Min. 1st Qu. Median Mean 3rd Qu. Max.
3.9 20.6 30.0 37.4 50.3 101.1
Another useful way to
visualize the spread of breast cancer data is through a boxplot (I will make it horizontal for easier comparison with the histogram):
> boxplot(gapCleaned$breastcancer, col=”blue”, horizontal=T, border=”red”, main=”Boxplot of breast cancer”, xlab=”Brest cancer new cases globally”)
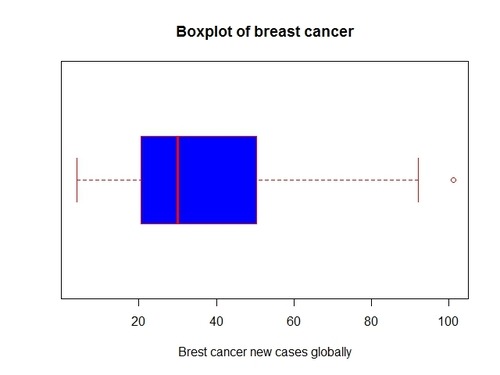
Note that the median is in the left half of the box and that the right whisker is longer that the right whisker, as typical for right-skewed distributions. The boxplot confirms our considerations made previously on the data.
Another interesting analysis on breast cancer, might also be
producing a frequency table of the data. However breast cancer values are pretty unique in each single country.
> freq (as.ordered(gapCleaned$breastcancer)) # print a frequency table
So, by now we should have a good understanding of distribution, spread and shape of breast cancer data at a global level. What I am going to do next is:
- First, see which countries present highest number of breast cancer new cases, and which ones the lowest
- Second, see if there is any significant difference in breast cancer between different continents. This can help us identifying hidden patterns in the data and potential associations between breast cancer and specific socio-economic aspects of individual countries/continents (I will go deeper on this with another post!)
Here
I sort countries by breast cancer to see most/least affected. As an example, I print the top and worst 5 countries.
> ordered<-gapCleaned[order(gapCleaned$breastcancer),c(1,5,17)]
> ordered[c(1:5, 169:173),] # top 5 vs worst 5
ID country breastcancer continent
165 Mozambique 3.9 AF
105 Haiti 4.4 LATAM
90 Gambia 6.4 AF
161 Mongolia 6.6 AS
206 Rwanda 8.8 AF
117 Israel 90.8 AF
86 France 91.9 WE
174 New Zealand 91.9 OC
23 Belgium 92.0 WE
269 United States 101.1 NORAM
Now
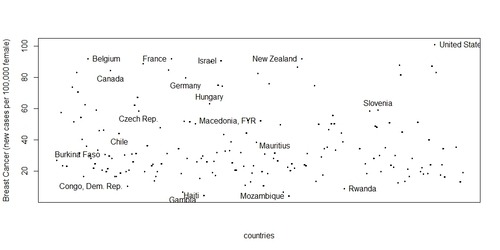
let see countries performance in a plot and identify again some of the most/least affected by breast cancer. (i don’t think this is a very nice way to visualize all countries performance but that is what I’ve been able to come up with for now! I would really like to plot breast cancer on a world map…I know this can be done with R and would probably need latitute/longitude coordinates, so if you can help me, please feel free to comment here)
> plot(gapCleaned$country, gapCleaned$breastcancer, xaxt=”n”, xlab=”countries”,ylab=”Breast Cancer (new cases per 100,000 female)”)
> identify(ordered, labels=ordered$country)
Let see now
breast cancer by continent: are there significant differences? Note in the original dataset there was no variable indicating the continent of the country. So I added myself a categorical variable including 7 levels (continents) as below:
> unique(gapCleaned$continent) # print levels of my categorical variable continent
[1] AS EE AF LATAM OC WE NORAM
We can also see how many countries we have in the data set for each continent.
> table(gapCleaned$continent)
AF AS EE LATAM NORAM OC WE
0 56 35 22 28 3 8 21
Finally lets plot breast cancer by continents.
To plot relationship between a categorical variable (continent) and a quantitative variable (breast cancer), we can use boxplots. Are there significant differences between continents?
> boxplot(gapCleaned$breastcancer ~ gapCleaned$continent, main=”Breast cancer by continent”, xlab=”continents”, ylab=”new cases per 100,00 residents”, col=gapCleaned$continent)
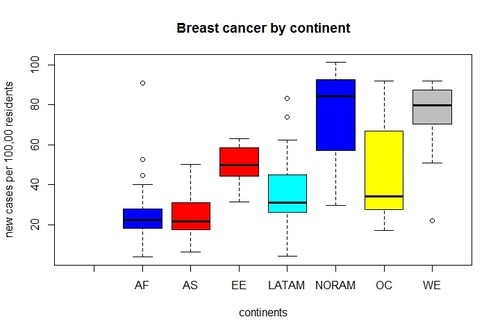
Yes.
It looks there are significant variations in breast cancer among continents. We can clearly see that the median of breast cancer for North America and West Europe is substantially higher than other continents. On the other hand, Africa and Asia report the lowest number of new cases of breast cancer.
This let me immediately think whether there might be a positive correlation between economic development of a country (more specifically GDP per capita) and inclination to breast cancer. Are women living in rich countries more likely to contract breast cancer? To be honest,
I don’t really think there is a strong direct relationship between the two variables, much less a causation. Yes, richer countries might be associated with higher cases of breast cancer detected; but at the end of the day it depends on how this wealth is spent by people… is the money spent to buy high-calorie foods or alcoholic drinks that in the long run will put your health at risk? or is it invested to manufacture goods that generate more CO2 emissions? I will definitely explore if there is any association between breast cancer and wealthy of a country, but as initially mentioned, my research question will focus on people lifestyle and specifically whether women conducting a healthy/unhealthy lifestyle are less/more likely to contract breast cancer.
Only data can give me an answer ;)
see you next post!